
A Novel Biofoundry Blueprint
Published in Nature Communications, the international consortium’s four-tier abstraction hierarchy standardises every design-build-test-learn step, opening the door to fully interoperable, AI-optimised biomanufacturing across the globe.
Glossary
DBTL - Design-Build-Test-Learn
SBOL - Synthetic Biology Open Language
LabOP - Laboratory Open Protocol Language
Main Text
A Nature Communications Perspective published on 1 July 2025 revealed a four-tier abstraction hierarchy that organises every biofoundry from high-level “Project” aims down to granular “Unit-operations”. Crafted by an international team spanning KRIBB, Lawrence Berkeley National Laboratory and Imperial College London, the framework gives the hardware and software a shared lexicon, so facilities in different countries can exchange protocols and data without translation frictions. Imperial College’s London Biofoundry (which co-authored the work) stressed that the schema directly tackles the interoperability bottleneck hindering collaboration across the 33-member Global Biofoundries Alliance [1].
The abstraction hierarchy has the following structure:
- Level 0 | Project - represents a series of tasks to fulfil the requirements of external users who wish to use the biofoundry.
- Level 1 | Service/Capability - refers to the functions that external users require from the biofoundry and/or that the biofoundry can provide.
- Level 2 | Workflow - refers to the DBTL-based sequence of tasks needed to deliver the Service/Capability. Each workflow is intentionally assigned to a single stage of the DBTL cycle to ensure modularity and clarity in execution.
- Level 3 | Unit-operations - represents the hardware or software that will perform the tasks required to fulfil the desired workflow (42 hardware and 37 software modules in the initial catalogue).
This tiered decomposition lets any foundry mix-and-match modules in silico and run them on-site with minimal human intervention.
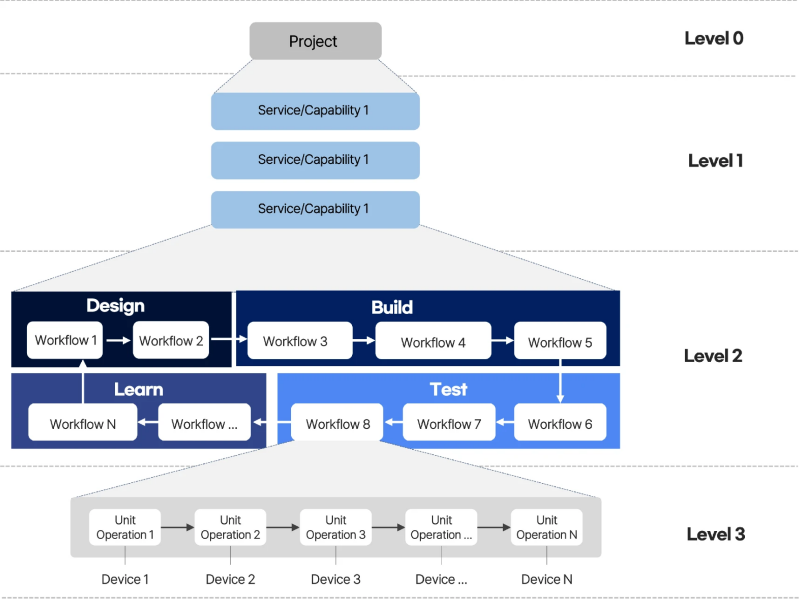
Abstraction hierarchy of biofoundry operations across four levels: project (lvl 0), service/capability (lvl 1), workflow (lvl 2) and unit operation (lvl 3). Source: Nature Communications
The difficulty arises with the diversity of biological experiments and the continuous development of improved equipment and software. The predefined workflows and unit-operations are meant as flexible templates: in practice a single label (e.g., “Liquid Media Cell Culture”) can cover very different experimental routines while laboratories oftentimes adapt, merge or split steps to suit their specific goals and equipment.
To make modular workflows genuinely interoperable, biofoundries must exchange protocols in a common data language: existing standards such as SBOL and LabOp already encode DBTL tasks and instruments, and SBOL’s mature tool-chain can ride seamlessly on the new abstraction hierarchy; the next step is to populate that scaffold with foundry-specific protocol libraries, recognising that many unit-operations simply repackage familiar methods (e.g., Golden Gate Assembly as a string of liquid-handling and thermocycling steps). By treating these as shareable, ontology-backed building blocks rather than lab-specific recipes, the framework paves the way for calibrants, measurands and ultimately reproducible, comparable operations across the global biofoundry network.
" ... workflows and operations can differ greatly between biofoundries. This highlights the need for open standards and methodologies to enable protocol exchange." - Professor Paul Freemont [2]
Biofoundries are becoming an evermore integral part of modern life science research. These state-of-the-art facilities collapse the traditional DBTL cycle into an automated, high-throughput pipeline that allows scientists to prototype thousands of genetic constructs in days rather than months. This speed translated directly into the rapid creation of mRNA COVID-19 vaccines and is now propelling antibiotic-discovery campaigns and patient-specific cell therapies. Automation removes much of the variability introduced by manual lab work, so results are more reproducible and can feed seamlessly into ML models that guide the next round of experiments, creating a virtuous feedback loop. Beyond accelerating research, biofoundries also lower costs and democratise access to biotechnology [3].
Overall, this development marks a decisive move toward a truly interoperable global biofoundry network - one that will streamline collaboration and accelerate synthetic biology research, equipping the field to tackle urgent scientific and societal challenges more swiftly.
References:
[1] Kim H, Hillson NJ, Cho B-K, Sung BH, Lee D-H, Kim D-M, et al. Abstraction hierarchy to define biofoundry workflows and operations for interoperable synthetic biology research and applications. Nature Communications 2025.
[2] Ntumba R. Pioneering paper lays groundwork for smarter, connected biofoundries worldwide. Imperial News 2025.
[3] Biofoundries: a revolution for research and global economy. Bioconvs.org


Add comment
Comments